Your body is comprised of roughly 37 trillion cells. If you peer inside any one of them, what you see would resemble a city: highways with tiny cargo, power plants releasing energy and elaborate farms that keep the cell fed. Making these cellular cities thrive requires a special kind of protein that physically links different parts of the cell together. Copies of this protein, placed at hundreds of locations, serve as a decentralized communications network — a high-speed internet of miniature proportions.
This protein — fundamental to the life of a cell and, thus, life — should exist in every cell of every organism on the planet. However, despite scientists’ best efforts, it had been identified only in primitive yeast cells, and nowhere else. Some had begun to doubt it was as ubiquitous as initially thought. But now, the search is over.
In a study published today in Science, researchers at Columbia’s Mortimer B. Zuckerman Mind Brain Behavior Institute identified the mystery protein, called PDZD8, in the cells of mice and humans.
We spoke with Franck Polleux, PhD, Zuckerman Institute Principal Investigator and the paper’s senior author, about how his lab found PDZD8, and how this research stands to usher in a new era of discovery — where the smallest parts of the cell can be used to tease apart biology’s largest questions.
This research stands to advance our understanding of how cells function and, as a result, the cellular basis of the most complex and persistent diseases we face.
Why was it so important to identify this protein?
This protein represents one of the most fundamental components of biological life. Just as your body is made up of organs — your liver, your stomach, your heart and so on — each individual cell has organs of its own, called organelles, which work in harmony to keep the cell healthy. Organelles communicate by physically connecting, or ‘tethering,’ to each other.
In our study, we investigated the protein that tethered together two organelles: the mitochondria, which serve as the cell’s main energy source, and the endoplasmic reticulum (ER), a cellular factory that churns out proteins and ships them to other parts of the cell.
The ER tether to mitochondria at hundreds of locations. But visualizing those tether sites was virtually impossible; a fundamental feature of cells that no one could truly study. Scientific questions remained unanswered, lines of inquiry lay stagnant or were abandoned.
With these new findings, we hope scientists will revisit old investigations, or launch entirely new ones. This research stands to advance our understanding of how cells function and, as a result, the cellular basis of the most complex and persistent diseases we face — from Alzheimer’s disease to diabetes to cancer.
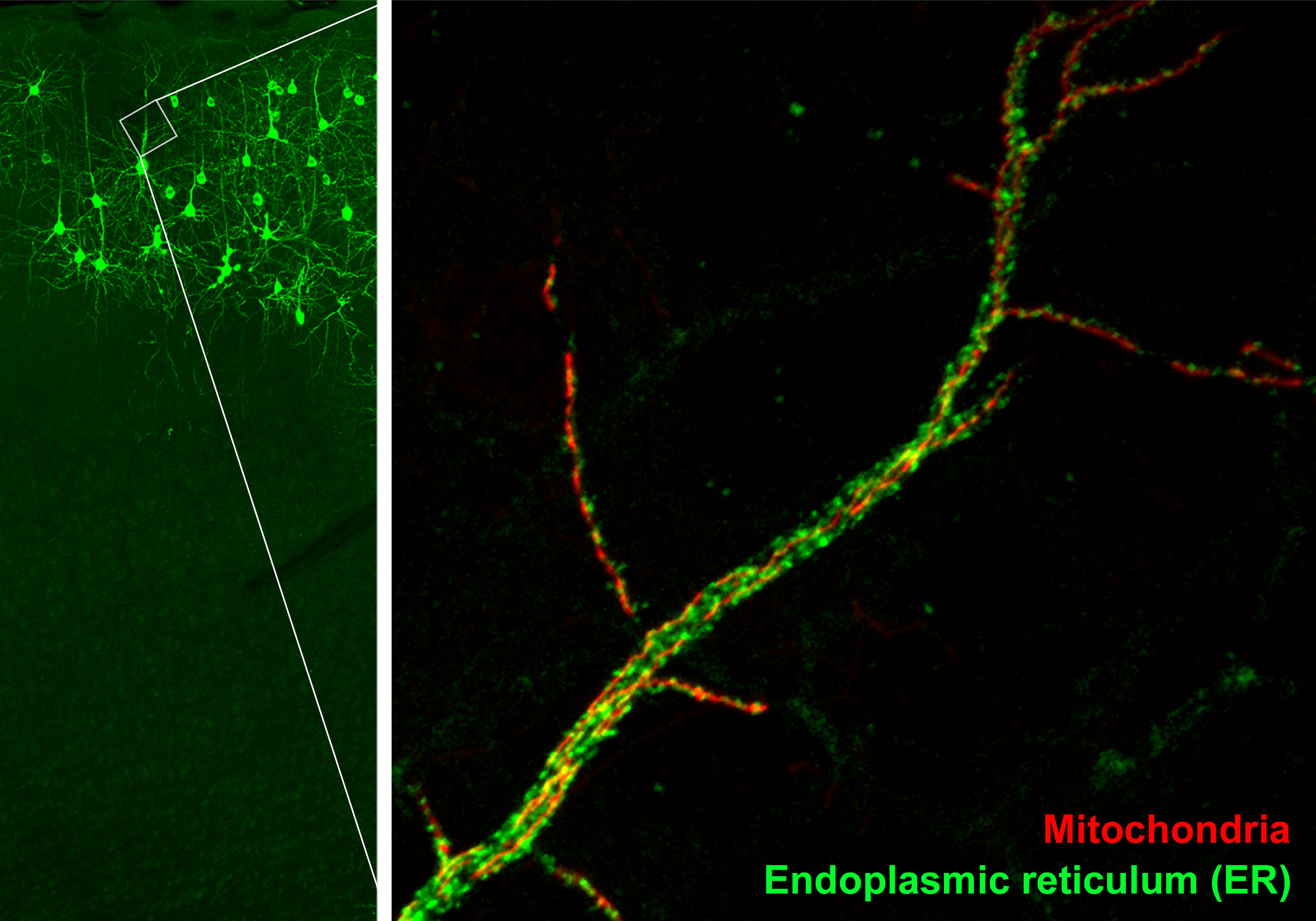
Zoomed-in image of mitochondria (red) and endoplasmic reticulum (green) in the dendrite of a neuron (Credit: Yusuke Hirabayashi/Polleux Lab/Columbia's Zuckerman Institute).
What was your starting point for finding this protein?
Most of what we know about ER-mitochondria tethering has come from studies in baker’s yeast, a single-celled organism that shares many features with cells of more complex multi-celled organisms, including humans.
Eight years ago, Dr. Peter Walter and his lab at the University of California at San Francisco identified in yeast a cluster of four proteins, called the ERMES complex, which appeared to physically tether ER to mitochondria. Bolstered by this discovery, scientists began looking for its equivalent— known as an ‘ortholog’ — in animals. But for several years, no one could find it; not until we employed an entirely different method of discovery.
This was our first big breakthrough, and set in motion an ambitious investigation that, over the next four years, helped us decipher the purpose of this elusive protein.
Normally when comparing distantly related proteins, scientists examine what each protein is made of; the more similar the proteins’ components are to each other, the more likely they perform the same function. However, this is a poor way to predict similarities between yeast and mammals, which are separated by 700 million years of evolution.
But recent advances in structural biology and computation now allow scientists to identify similarities by comparing proteins’ physical structure. So, by searching for proteins in mammalian cells that had a similar structure to those of the ERMES complex, we identified about half a dozen potential orthologs.
With multiple candidate proteins, how did you narrow it down?
This was challenging for a relatively small lab like mine. But because we knew our mystery protein tethered ER and mitochondria, we reasoned it would be located on the outer surface of one of those organelles.
Applying that criteria to the candidates left us with just two proteins, one of which was PDZD8. We then used CRISPR-Cas9, a powerful gene-editing technique, to tag PDZD8’s genetic code with a small fluorescent protein and confirm its location on the surface of ER, specifically at ER-mitochondria connection sites.
What did you do next?
If PDZD8 worked like its yeast equivalent, we should be able to swap it into yeast cells and have it perform in nearly the same way. But we knew this would be challenging because of the long evolutionary gap between yeast and mammals.
But a talented graduate student named Wolfgang Pernice from Dr. Liza Pon’s lab at Columbia performed the swap, confirming that PDZD8 served largely the same function as ERMES. Now we needed to see PDZD8 in action in mammalian cells. And because cells are complex, three-dimensional physical entities, we needed to see them in 3D.
How is it possible to image something so small?
The space between ER and mitochondria is just 30 nanometers — 3,000 times smaller than the diameter of a human hair. But we had powerful imaging technology at our disposal called a Focused Ion Beam-Scanning Electron Microscope. This microscope uses an electron beam to image the surface of a block of tissue. Then a diamond blade shaves off 10 nanometer-thin slices. After shaving the first slice, the microscope then images again, then slices again, repeating this cycle thousands of times. This allows us to see entire cells at incredibly high resolutions (down to the level of individual molecules). We first imaged normal human cells, and then in cells with deactivated PDZD8.
From there we painstakingly reconstructed these image slices into a 3D model. We then confirmed that switching off PDZD8 completely untethered ER from mitochondria, but everything else — including the underlying structure of both organelles — remained intact.
This was the moment when we realized PDZD8 was what the scientific community had been searching for.
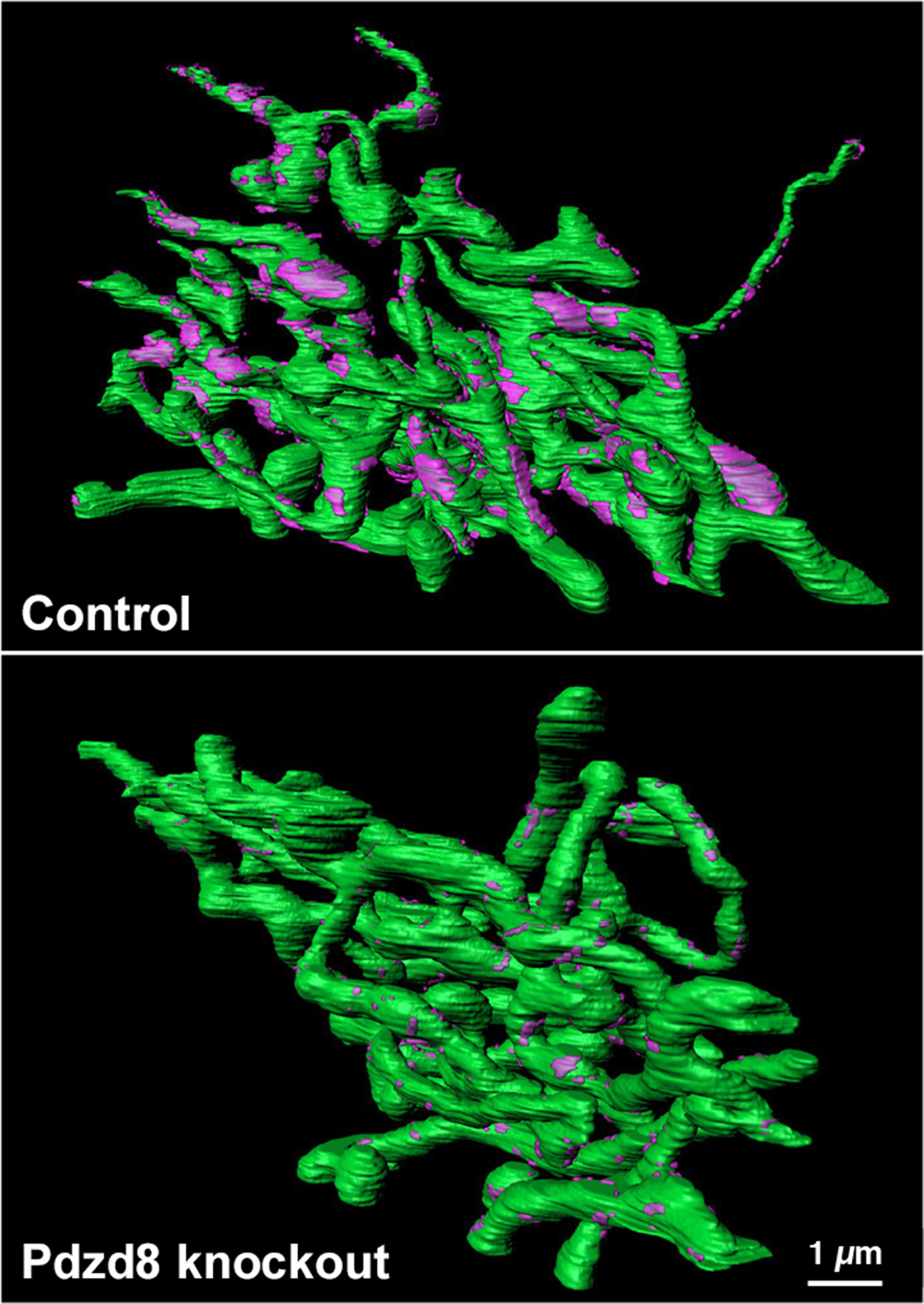
Reconstruction of mitochondrial-ER contact points in a control cell (top) compared to a cell lacking PDZD8 Credit: Yusuke Hirabayashi/Polleux Lab/Columbia's Zuckerman Institute)
What did you discover about the function of this protein?
After performing additional experiments in both mouse and human cells, including neurons, we confirmed that PDZD8 mediates communication between organelles by sending calcium ions, which act as messenger molecules, from the ER to mitochondria. Disruptions to this communication can be harmful to the cell, and points to the fundamental importance of PDZD8.
Inside ER are stores of calcium ions. The prevailing wisdom was that ER communicates with mitochondria by releasing the calcium ions diffusely throughout the cell. But we found that by tethering ER to mitochondria, PDZD8 allows ER to send calcium ions directly into the mitochondria. Very little seeps out into rest of the cell. And in neurons that lack PDZD8, calcium ions are more likely to end up floating around in the cell, instead of being transferred directly into mitochondria, causing a disruption in the cellular communications network that could play a factor in the cellular basis of a variety of diseases and disorders.
As a neuroscientist, how does this study advance efforts to understand the brain?
This discovery has opened up an entirely new line of investigation into how neurons use calcium ions to communicate, whether it’s between different organelles inside the same neuron, or among several neighboring neurons. As we decipher the mechanics of this process, we hope to understand how neurons and groups of neurons function.
Moreover, recent studies have shown that disruptions to calcium-ion flow may play a role in diseases such as Alzheimer’s, which is of particular interest in my lab. So many areas of inquiry were stalled because this protein couldn’t be found. Now we have a new, powerful tool to understand the underlying causes of — and move closer to treatments for — this and other complex and devastating brain diseases.
###
This paper is titled: “ER-mitochondria tethering by PDZD8 regulates Ca2+ dynamics in mammalian neurons.” Additional contributors include co-first authors Yusuke Hirabayashi, PhD, and Seok-Kyu Kwon, PhD as well as Hunki Paek, PhD, Wolfgang Pernice, PhD, Maëla Paul, Jinoh Lee, Parsa Erfani, Ashleigh Raczkowski, Donald Petrey and Liza Pon, PhD.
This research was supported by the National Institutes of Health (NS089456, GM122589, GM45735, S10 RR02300, 1S10OD016167), the Eunice Kennedy Shriver National Institute of Child Health & Human Development (U54 HD083092), the Fondation Roger De Spoelberch, the Japan Society for the Promotion of Science, JST/PRESTO (JPMJPR16F7) and the Simons Foundation.
The authors report no financial or other conflicts of interest.