Every animal, from an ant to a human, contains in their genome pieces of DNA called Hox genes. Architects of the body, these genes are keepers of the body’s blueprints; they dictate how embryos grown into adults, including where a developing animal puts its head, legs and other body parts. Scientists have long searched for ways to decipher how Hox genes create this body map; a key to decoding how we build our bodies.
Now an international group of researchers from Columbia University and the Spanish National Research Council (CSIC) based at the Universidad Pablo de Olavide in Seville, Spain have found one such key: a method that can systematically identify the role each Hox gene plays in a developing fruit fly. Their results, reported recently in Nature Communications, offer a new path forward for researchers hoping to make sense of a process that is equal parts chaotic and precise, and that is critical to understanding not only growth and development but also aging and disease.
“The genome, which contains thousands of genes and millions of letters of DNA, is the most complicated code ever written,” said Richard Mann, PhD, principal investigator at Columbia’s Mortimer B. Zuckerman Mind Brain Behavior Institute and the paper’s co-senior author. “Deciphering this code has proven so difficult because evolution wrote it in fits and starts over hundreds of millions of years. Today’s study offers a key to cracking that code, bringing us closer than ever to understanding how Hox genes build a healthy body, or how this process gets disrupted in disease.”
We now had a starting point from which to systematically decode Hox gene regulation; a kind of Rosetta Stone to help us decipher the genetics of body development.
Hox genes are ancient; they can be found across all animal species. Even primitive jellyfish have them. Each type of organism has different combinations of these genes. Fruit flies have eight Hox genes, while humans have 39.
These genes work by producing special proteins called transcription factors, which work together with similar proteins called Hox cofactors to bind to different segments of DNA and turn many other genes on and off at just the right time — a Rube Goldberg machine of microscopic proportions.
“Because these genes are intricately involved in many aspects of development, it has proven incredibly challenging to isolate individual Hox genes and trace their activity over time,” said James Castelli-Gair Hombría, PhD, a principal investigator at the Centro Andaluz de Biología del Desarrollo at the Universidad Pablo de Olavide and the paper’s co-senior author. “We had this incredibly complex equation to solve but too many unknowns to make significant progress.”
Recently, Dr. Hombría and his team hit upon a bit of luck. While examining genetic activity in a developing fruit fly, they stumbled upon a small piece of regulatory DNA, called vvI1+2, that had an unusual — and surprising — attribute. Although it was active in cells across the fruit fly’s entire developing body, it appeared to be regulated by all of the fruit fly’s eight Hox genes.
“The ubiquity of the vvI1+2 DNA segment across the entire developing fruit fly, combined with the fact that every Hox gene touches it, made it an ideal system by which to study the Hox gene family,” said Carlos Sánchez-Higueras, PhD, a postdoctoral researcher in the Hombría lab and the paper’s first author. “In this single piece of DNA, we had the perfect tool; we could now devise a method to systematically manipulate vvI1+2 activity to see how each Hox gene functioned.”
First, Dr. Sánchez-Higueras teamed up with Dr. Mann at Columbia’s Zuckerman Institute and used a sophisticated computer algorithm called No Read Left Behind, or NRLB. NRLB was recently developed by Dr. Mann, his lab, and his collaborators, including Columbia systems biology professor Harmen Bussemaker, PhD. This powerful algorithm pinpoints the locations where transcription factors bind to a stretch of DNA, even if these binding sites are very weak and difficult to capture. For this study, the researchers focused on the Hox transcription factors and Hox cofactors that bind to vvI1+2.
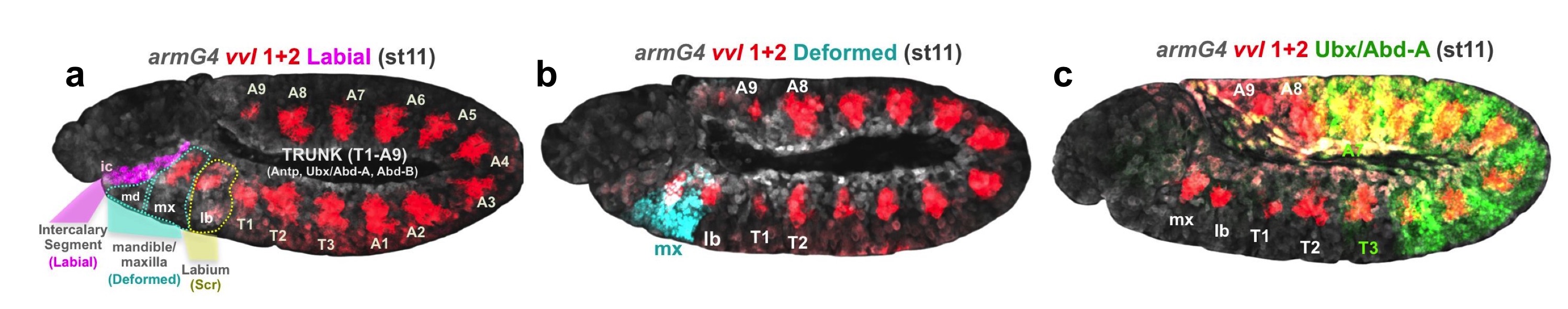
Drosophila melanogaster developing embryos showing the spatial relation of vvl1+2 expression pattern (red) with respect to various Hox expression domains. a Labial (purple) is expressed in the intercalary segment, (b) Dfd (blue) is expressed in the mandible and maxilla, (c) Ubx and Abd-A (green) protein expression extends from T3 to A7 (Credit: Carlos Sanchez-Higueras/Hombría Lab/CABD).
“Our analyses provided a precise road map of Hox binding sites in vvI1+2, which we could then apply to a living fruit fly,” said Dr. Mann, who is also the Higgins Professor of Biochemistry and Molecular Biophysics (in Systems Biology) at Columbia’s Vagelos College of Physicians and Surgeons.
By employing a combination of elegant genetic manipulations in living, or in vivo, fly embryos, together with advanced biochemical and computational analysis, the researchers could then systematically manipulate Hox target activity with an unprecedented level of precision.
“We now had a starting point from which to systematically decode Hox gene regulation,” said Dr. Hombría, “a kind of Rosetta Stone to help us decipher the genetics of body development.”
The researchers’ findings are especially promising because they can be applied to the entire genome. The steps that Hox genes undergo to regulate vvI1+2 can inform how Hox genes regulate other DNA, not just in fruit flies but beyond — including in vertebrates, such as mammals, and even humans.
“While there is much about Hox genes that remains to be elucidated, our work is a significant step forward,” said Dr. Sánchez-Higueras. “These continued efforts, combined with those of our peers, will help shed light on how the whole system works together in a growing embryo and how they contribute to disease.”
###
This paper is titled “In vivo Hox binding specificity revealed by systematic changes to a single cis regulatory module.” Additional co-authors include Chaitanya Rastogi, PhD, Roumen Voutev, PhD, and Harmen Bussemaker, PhD.
This research was supported by the María de Maeztu Unit of Excellence, the Ministerio de Ciencia e Innovación, the European Regional Development Fund, the European Molecular Biology Organization and the National Institutes of Health (R35 GM118336, R01HG003008).
The authors report no financial or other conflicts of interest.